Cell-based immunotherapy has gained great attention from researchers and pharmaceutical companies, particularly due to its promise to treat various cancers. Understanding the interaction between drug candidates and targets on the cell membrane is crucial to drug development. One area of particular importance is the interaction between drugs and immune cells.However, this is a complex biological process, which not only depends on the drug molecules and receptors engaged, but also on the possible clustering and dimerization of the receptors induced by the molecular binding. Techniques used for studying the interaction between drug molecules and their targets often require the target proteins to be extracted from the cell or the cells being fixed. Unfortunately, the integrity of a live cell could be compromised by protein extraction and cell fixation. Therefore, it is advantageous to directly characterize the molecular interaction of living cells.
SPR microscopy (SPRM) has been demonstrated as a viable technique capable of measuring binding affinity and kinetics at cell membranes. We demonstrate in this application note that SPRM can perform kinetic studies of living cells. Compared to instruments designed for studies of live cells, such as FACS and LigandTracer, SPRM is a label-free technique for measuring binding kinetics and affinity in real time. The system integrates SPR and optical microscopy for simultaneous cell imaging and SPR binding measurements.
In here, we studied the binding of Rituximab, a mouse/human chimeric monoclonal bivalent antibody (Fig. 1), to the CD20 receptors, which are endogenously expressed by B cells. This antibody was approved by the US FDA in 1997 to treat B-cell non-Hodgkin lymphomas. It acts by depleting normal as well as pathogenic B cells while sparing plasma cells and hematopoietic stem cells, which do not express the CD20 surface antigen.
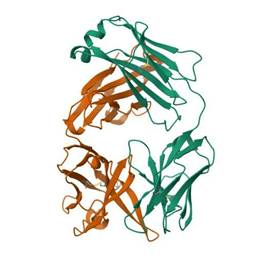
Fig. 1: Structure of rituximab Fab (pdb_00004kaq)
Rituximab was purchased from InvivoGen. Ramos cells, purchased from ATCC, were cultured in a growth medium before being transferred to about 1 mL of imaging buffer at a density of about 1 x 106 cells/mL. This cell imaging buffer was also used as the running buffer. Au sensor chips coated with PLL were used as the substrate for capturing and anchoring B cells. About 50 uL Ramos cell solution was pipetted onto the chip. Capture of the Ramos cells was simultaneously monitored by the bright-field camera and SPRM. Additional aliquots of the Ramos cell solution can be added onto the chip if a greater confluence is desired. Each chip was washed with a running buffer to remove excessive cells before mounting onto the SPRM. Seven serial dilutions were made from a 75 nM stock in the running buffer (x3 dilution) and injected into the system using the kinetic titration method.
Captured Ramos cells were stable on the PLL coated chip surface during the experiment. As seen in the bright field image in (Fig.2a), they maintained their spherical shape with no obvious degradation. The statistical analysis of binding affinity and kinetic constants reveals the heterogeneity between cells and the averaged kinetic constants. The red squares on the cells indicate binding activity for the statistical calculation of the kinetic parameters, which overlap well with the actual positions of the attached cells.
High resolution SPRM measurements were generated by segmenting the sensing area into 2400 regions of interest (ROI). For every ROI, a 1:2 heterogeneous kinetic binding interaction model was applied. ROIs that detected a response which fitted well to the kinetic interaction model were collectively plotted onto an iso-affinity scatter plot for statistical analysis of the binding heterogeneity and two kinetic modes of bi-valent interactions. Histograms extracted from the iso-affinity scatter plot were fit with Gaussian distributions to obtain mean values and standard deviations for the association rate (ka), dissociation rate (kd), and equilibrium dissociation constant (KD) for both the modes of interaction (Fig.2c). The average equilibrium and kinetic constants are KD = 3.5 nM, 424 nM, ka = 8.5 x 104 M-1 s-1 , 1.3 x 104 M-1 s-1 and kd = 1.6 x 10-4 s-1, 2.9 x 10-2 s-1. The statistical distribution of these values reflects the dynamic complexity of the living system that affects the binding process.

Fig. 2. A) Bright field and B) SPR images of Ramos cells. Red squares show valid binding overlapped with cells, C) Association rate constants ka, dissociation rate constant kd, and Affinity KD measured for Rituximab binding CD20 receptors expressed on living B cells.
In summary, SPRM can measure binding affinity and kinetic constants of membrane targets at living cells, which are difficult to obtain using traditionally label-based and label-free methods. Measuring biophysical properties of a target in the native membrane environment in real time without labels provides more accurate information about the drug-receptor interactions, opening an opportunity to speed up drug development, particularly for molecularly targeted therapy.
We thank Nanxi Yu from ASU for assisting with the samples and Jon Brooks from Pfizer for the valuable discussions.
DOWNLOAD PDF
Download a PDF of Application Note 141: SPRM Measurements of Binding Kinetics between Rituximab and CD20 on Live B Cells
