As the coronavirus 2019 (COVID-19) pandemic continues to cause sever impacts on human life and world economy, scientists are rushing in to researches to understand the disease. This disease is caused by the highly contagious coronavirus 2 (SARS-CoV-2). During viral infection, the trimeric COVID-19 spike-protein (S-protein) is cleaved into S1 and S2 subunits, where the S1 subunit contains the receptor binding domain (RBD), which directly binds to the peptidase domain of angiotensin (Ang)-converting enzyme 2 (ACE2) (1,2). Therefore, it is important to understand the binding interaction between S1 and S RBD proteins with ACE2, and Surface Plasmon Resonance (SPR) can play an important role in kinetic measurements of binding between ACE2 and S1 proteins to help address this worldwide health crisis.
In the conventional SPR approach, a series of analyte concentrations are injected into a fluidic channel pre-immobilized with a specific ligand density. A regeneration step is necessary to recover the ligand before a new analysis. This is accomplished by introducing a reagent such as acid, base, surfactant, or other compounds to dissociate the preformed bioconjugates at the surface. However, exposure to such harsh reagents can lead to irreversible conformational changes of the immobilized ligand or receptor, which might not recover even after being allowed to renature in a buffer for an extended period. Moreover, in many cases the preformed bioconjugates are so tightly bound that a suitable regeneration solution cannot be found. Thus, the reliability of the SPR data can be seriously compromised as the “regeneration” step is not reliable or simply cannot be performed. Even with ligands that can be completely recovered, injections of multiple samples upon repeated regenerations are both time and sample consuming.
This application note shows the application of the single-injection measurement technique in binding kinetic studies between COVID-19 S1 and ACE2 proteins. The five-channel SPR instrument, with the BI-DirectFlow™ technology, provides a unique multi-channel technique for precise sample delivery and high-quality measurement. Variable ligand densities can be controllably formed with a single injection of ligand solution. With a subsequent injection of a single analyte sample solution, four SPR sensorgrams can be simultaneously collected to determine the kinetic constants of the binding (3). The advantages of varying ligand densities include measurement of biomolecular binding that is impossible to accomplish with surface regeneration, simultaneous attainment of binding equilibria in all fluidic channels, rapid validation of the method without any ambiguity associated with secondary effects, and reduction in consuming expensive or precious biological specimen.
The biotinylated ACE2 protein was diluted in the running buffer and immobilized onto a streptavidin (SA) sensor chip using BI’s automated graduated injection technique, which immobilizes different amounts of ligand in each channel with a single injection. The ligand immobilization occurred in channels 1, 2, 3, and 4 for 200, 50, 15, and 5 s, respectively, affording a range of ligand densities. Figure 1 shows the responses from different channels where the immobilization densities of ACE2 can be deduced from the net change in baseline.
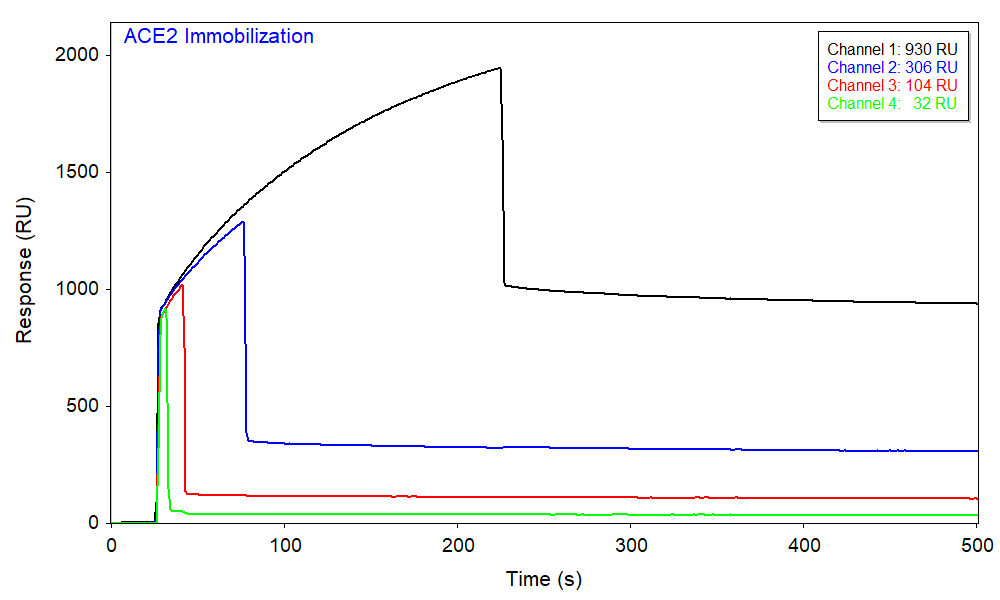
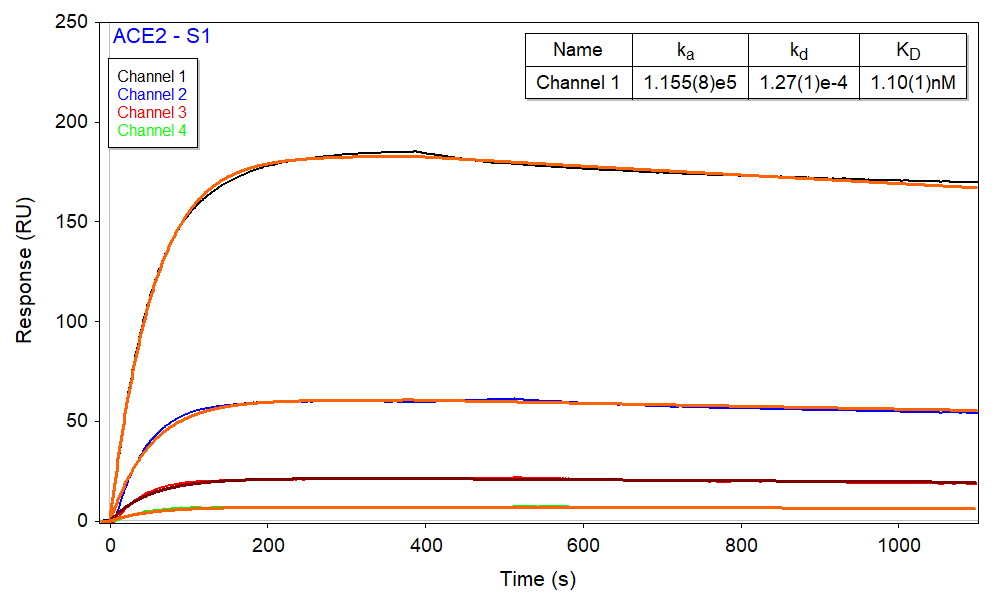
Afterwards, a single aliquot of 170 nM S1 was injected to all 5 channels with different ligand densities, where channel 5 has no ACE2 (i.e. zero ligand density) and serves as a reference. As expected, based upon the relative density of immobilized ACE2 in each channel, channel 1 produced the largest binding response and channel 4 produced the smallest (Figure 2). The sensorgrams can be fitted with the kinetic 1:1 interaction model. Kinetic parameters were obtained from a global fitting of all four channels with different ACE2 immobilization densities, as shown in Figure 2. The results agree well with the literature values and the SPR kinetic-titration measurement performed separately and are summarized in the Table 1 below.
| Parameter | ka [1/M*s] | kd [1/s] | KD [nM] | * EC50 [nM] | **KD by BLI [nM] |
| Single-injection | 1.16×105 | 1.27×10-4 | 1.10 | 5.99 | 3.95 |
| *** Kinetic-Titration | 1.85×105 | 2.15×10-4 | 1.10 | 5.99 | 3.95 |
* EC50 is 0.605 μg/ml from FACS analysis of SARS-CoV-2 S1 protein, Mouse IgG2a Fc Tag (Cat. No. S1N-C5257) binding to Vero E6 cells surface ACE2.
** KD determined for SARS-CoV-2 S1 protein, Mouse IgG2a Fc Tag (Cat. No. S1N-C5257) binding to Human ACE2, His Tag (Cat. No. AC2-H52H8) in BLI assay.
*** BI Application Note 138.
This note demonstrates the capability of measuring the binding interaction of SARS-CoV-2 viral protein (S1) with ACE2 using a multi-channel, single-injection method, a much simplified SPR procedure that saves time and reduces sample consumption.
DOWNLOAD PDF
Download a PDF of Application Note 139: Single-Injection Kinetic Measurement Technique for Studies of COVID-19 S1 Protein and ACE2 Binding
- Abassi Z et al., Front. Physiol. 2020,11, 574753.
- Renhong Yan et al., Science 2020, 367, 1444–1448.
- Xiaoying Wang et al., Anal. Chem. 2017, 89, 6, 3261–3265.
