Acid-sensing ion channels (ASICs) are voltage independent cation channels, which are expressed in both central and peripheral neurons.[1] Four genes that encode six ASIC subunits have been identified in mammals (ASIC1a, ASIC1b, ASIC2a, ASIC2b, ASIC3 and ASIC4) and these ASIC subunits assemble into homo and hetero-trimmers with singular biophysical properties.[2] ASICs are important drug targets involved in nervous (e.g., pain and diabetic neuropathy) and the central nervous systems (e.g., stroke, depression, neurodegenerative diseases). [3]
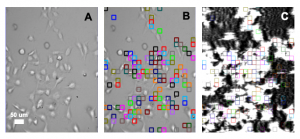
This application note describes the measurement of binding affinity (KD) of two antiparasitic diarylamidines (hydroxystilbamidine and pentamidine) to the ASIC1a receptor on HEK 293A cells with SPR Microscopy. SPR Microscopy is a technology that seamlessly combines both optical bright-field and SPR capabilities into a single system to enable label-free measurement of binding processes on single cells. The bright field provides topographical and phenotypical responses of the cells to a ligand, and the SPR measures the binding kinetics of the ligand to the receptors expressed on the cell surfaces.
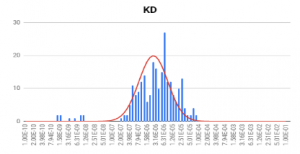
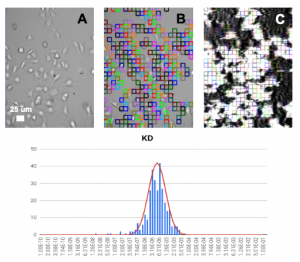
HEK 293A cells were grown and incubated on a SPR sensor chip for 24-36h, and the two ligands were introduced sequentially to binding to the ASIC1a receptor on each cell. Five concentrations of each ligand were injected into the system. Kinetic titration method was used to avoid the need of receptors regeneration after each ligand injection. KDvalues from ~600 regions (ROIs) were determined using the Langmuir binding isotherm model to provide unbiased and statistical information of the binding affinity. The measured mean KD are 2.6 uM for hydroxystilbamidine and 5.0 uM for pentamidine, which are consistent with literature data. [2,4] However, the SPRm 200 capability reveals statistical distribution in the binding affinity, which is associated with heterogeneity in the cells. This information cannot be obtained with other label-free affinity measurement techniques.
In conclusion, SPR microscopy allows study of binding affinity on the whole cells directly without using labels or lengthy sample preparations. Using SPRm 200, quantitative mapping of binding affinity of many hundreds of regions can be performed unbiasedly with unique statistical information on the heterogeneity of cells.
Author: Nguyen Ly | Biosensing Instrument | Published May 4, 2020
DOWNLOAD PDF
Download a PDF of Application Note 128: SPR Microscopy for Acid-Sensing Ion Channels
- Lingueglia, E et al, WIREs Membr Transp Signal 2013, 2:155-171
- Krauson AJ et al, PLoS ONE 13(5): e0196894
- Radu, B et al, Advances in Protein Chemistry and Structural Biology, 2016, 103: 137-167
- Chen, X et al, Neuropharmacology 2010 June, 58(7)