SPR microscopy (SPRm) has emerged as a unique tool for measuring the affinity and kinetics of ligand binding to membrane proteins on the cells directly. The technology integrates the traditional SPR and optical microscopy into one platform, which resolves individual cells spatially and quantity ligand binding to each of them in real time and without labels.
In this application note, we present small molecule compounds binding to a membrane protein overexpressed on human embryonic kidney cells, HEK 293. 60k cells overexpressed with a target receptor were seeded on the SPRm sensor chip and grown using standard protocols. A negative control sensor chip was also prepared by seeding 60k not-overexpressed HEK cells on another SPRm sensor chip. 7 dilutions of two small molecule compounds (eg SM1 & SM2) were prepared from 20 uM in standard buffer (x3 dilutions) and injected into the system using the kinetic titration method. The experiments were performed using a buffer containing 10 mM PBS, pH 7.4, 0.1% BSA and 0.02% DMSO.
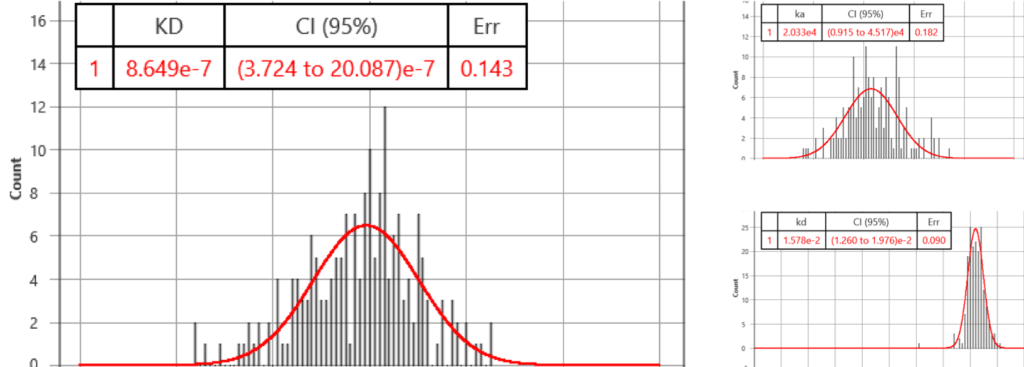
In the first run, compounds, SM1 and SM2, to the cells were sequentially injected into the system and the SPR signal of each cell to the compounds was measured. For SM1, the affinity (KD) and binding kinetic constants (ka and kd) were determined to be KD= 865 nM (372 – 2009 95% CI), ka= 2.0E4 (95%CI: 0.9 to 4.5) and kd= 1.6E-2 (95%CI: 1.3 to 2.0), respectively. These affinity and kinetic constants are consistent with functional assay results. SM2, a negative control compound, did not generate a SPR signal.
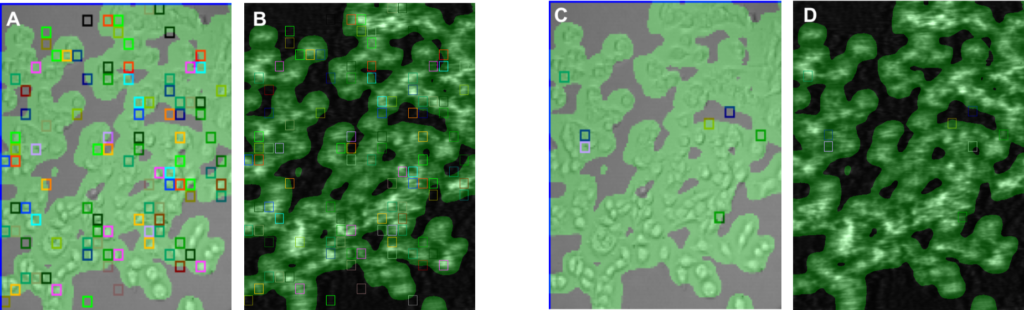

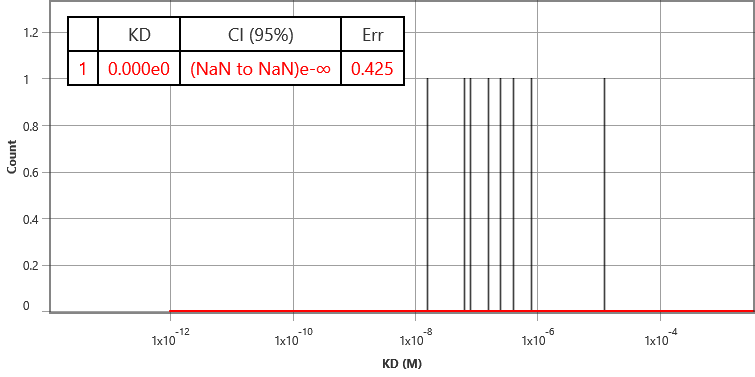
As a further control, binding of SM1 and SM2 to HEK cells that do not have the overexpssion of the receptors. None of the compounds showed any SPR binding signals, which confirms the specific binding of SM1 to the target membrane receptor.
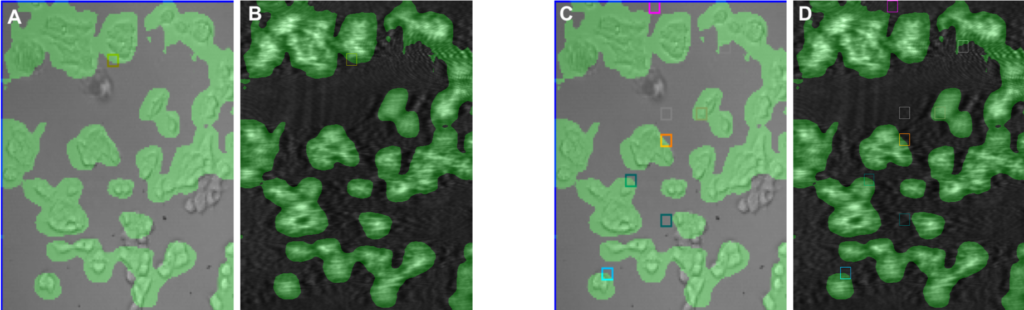
SPRm 200 is a powerful system that measures binding of small molecule to GPCRs directly on single cells without the need of extracting and purifying these receptors. The system provides real time biophysical analysis of binding events in a physiological setting.
We thank Genentech for the samples and useful discussions.
Author: Nguyen Ly | Biosensing Instrument | Published Jan 4, 2025
DOWNLOAD PDF
Download a PDF of Application Note 134: Positive and Negative Control Studies of Small Molecule Binding to Membrane Protein
