Antibody-drug conjugates (ADCs) combine the targeting specificity of monoclonal antibodies with potent cytotoxic payloads, enabling selective elimination of diseased cells while limiting systemic toxicity. As the clinical landscape of ADCs continues to grow, there is an increasing demand for analytical technologies that can directly study ADC interactions to receptors on intact cells.1 Surface plasmon resonance microscopy (SPRm) is a powerful, label-free, and real-time approach for resolving these interactions with high spatial and temporal resolution.2 In this application, SPRm implemented on a two-well sensor chip (Figure 1) was used to characterize the on-cell binding kinetics and specificity of two FDA-approved ADCs, a) Belantamab mafodotin and b) Gemtuzumab ozogamicin.
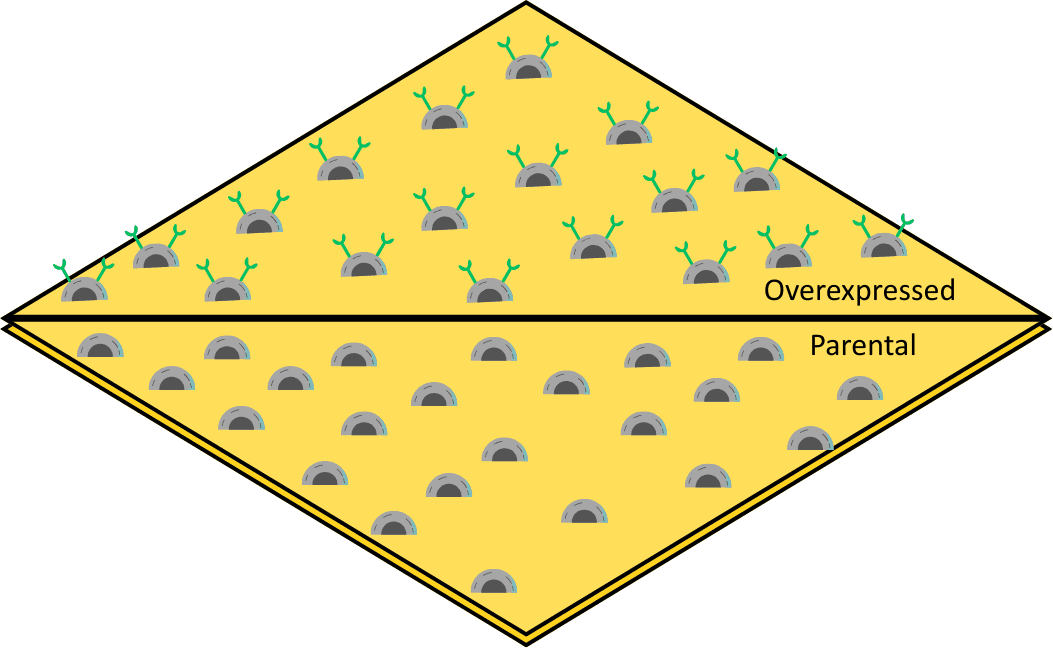
Figure 1: Schematic showing the SPRm setup with the two-well SPRm chip which has overexpressed U2OS cells on one side and parental U2OS cells seeded on the other side of the barrier.
Belantamab mafodotin is an IgG1-based ADC targeting B-cell maturation antigen (BCMA)and SPRm measurements revealed strong, high-affinity binding exclusively on BCMA expressing cells (Eurofins DiscoverX® PathHunter® U2OS BCMA Internalization Cell Line 93-1145C3), with no detectable interaction on parental U2OS cells not expressing the B-cell maturation antigen.3 Regions of interest (ROIs) were defined around individual cells from each population for analysis of the binding interaction. The red squares on the binding activity map reveal high amounts of cell-specific binding activity for the BCMA-ADC on U2OS cells expressing the target and the kinetic analysis revealed a distinct mode of binding, reflecting a simpler receptor engagement mechanism. Quantitative kinetic analysis showed a narrow distribution of KD values around the low nanomolar range (KD ≈ 3.4 × 10⁻⁹ M), which is typical for strong and homogeneous receptor binding. The isoaffinity scatter plot also indicates a tightly clustered population, which represents a relatively homogeneous binding process typical of monovalent binding (Figure 2). This clear discrimination between target-positive and target-negative cells confirms excellent on-cell specificity in a physiologically relevant context.
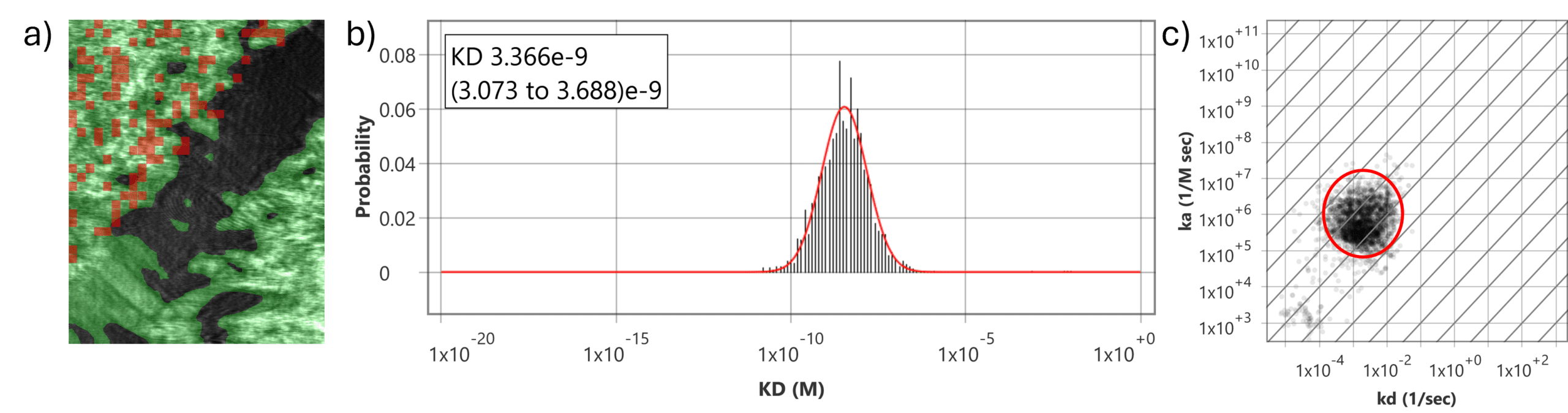
Figure 2: BCMA-ADC with overexpressed and parental U2OS cells. a) SPR image of a two-well SPRm chip with negative parental U2OS cells on top left corner and the positive U2OS cells overexpressing BCMA on the bottom right corner divided by a black barrier area. Red squares on the positive cell region indicate high cell specific activity. b) Histograms describing total kinetic interactions and distributions for BCMA-ADC with U2OS cells overexpressing BCMA receptors. c) Isoaffinity plot shows a corresponding prominent binding population. No kinetic data was obtained from the negative region containing the parental cells.
Gemtuzumab ozogamicin (Mylotarg), an IgG4-based ADC approved for CD33-positive acute myeloid leukemia, was also evaluated.4 Cells engineered to overexpress CD33 (Eurofins DiscoverX® PathHunter® U2OS BCMA Internalization Cell Line 93-1148C3) were measured alongside parental U2OS cells using the two-well chip. SPRm detected robust and selective binding of CD33-ADC on CD33 overexpressing cells, while no response was observed on parental cells, providing a stringent functional negative control. The KD histogram shows bivalent behavior, reflecting two affinity populations, and the isoaffinity scatter plot reveals two distinct kinetic populations (Figure 3). The initial interaction reflects the intrinsic monovalent binding affinity (KD ≈ 37 × 10⁻⁹ M) of a single Fab arm to CD33, while subsequent engagement of the second arm enhances overall binding strength (KD ≈ 6.7 × 10⁻⁹ M) through avidity, resulting in a more stable complex.
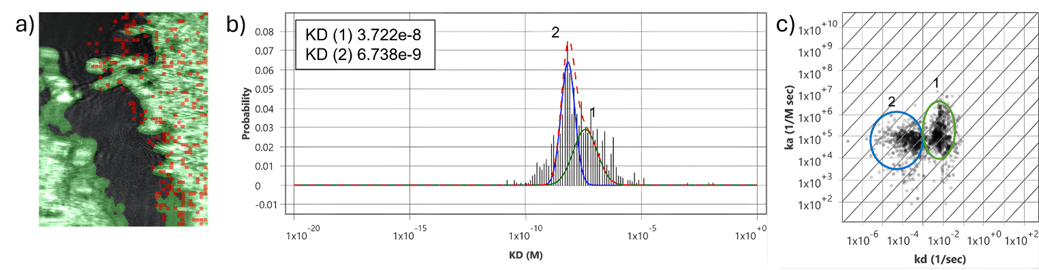
Figure 3: CD33-ADC with overexpressed and parental U2OS cells. a) SPR image of a two-well SPRm chip with the parental U2OS cells on the left and U2OS cells overexpressing CD33 on the right separated by a black barrier area. Red squares on the positive cell region indicate high cell specific activity on the positive cells. b) Histograms describing total kinetic interactions and distributions for CD33-ADC with U2OS cells overexpressing CD33 receptors. c) Isoaffinity plot shows two corresponding binding populations. No kinetic data was obtained from the negative region containing the parental U2OS cells.
The two-well SPRm chips enabled simultaneous, side-by-side analysis of target-positive and target-negative cells under identical experimental conditions, minimizing variability while improving experimental efficiency. Binding activity maps highlighted cell-specific ADC interactions at the single-cell level, while real-time kinetic traces provided quantitative insight into binding heterogeneity that is inaccessible using ensemble SPR methods. Together, these results demonstrate that SPRm is a sensitive and versatile platform for characterizing ADC binding kinetics directly on intact cells. By resolving receptor-specific interactions, monovalent binding, bivalent binding behavior, and cellular heterogeneity in real time, SPRm provides mechanistic insights that extend beyond traditional antibody assays. The integration of functional negative controls within a two-well chip further strengthens data confidence, making SPRm a valuable tool for ADC discovery, optimization, and translational cancer research.
We sincerely thank Eurofins DiscoverX for their collaboration on these measurements.
DOWNLOAD PDF
Download a PDF of Application Note 176: On-Cell Kinetic Characterization of Antibody–Drug Conjugates Using Two-Well SPR Microscopy
- Beck, Alain, et. al., Nature Reviews Drug Discovery 16.5 (2017): 315-337.
- Zhou, Xiao‐Li, et. al., Angewandte Chemie International Edition 59.5 (2020): 1776-1785.
- Barilà, Gregorio, et. al., Pharmaceuticals 14.1 (2021): 40.
- Sun, Te, et. al., Molecular and Clinical Oncology 18.6 (2023): 47.
