Incomplete synthesis of glycans or the overexpression of certain glycans promote a tumor-specific glyco-code.1 The glyco-code includes increased branched N-glycans, truncated O-glycans, hyper-sialylated glycans or glycolipids.2&3 Non-catalytic glycan binding proteins called lectins can recognize well-defined epitopes within a glycan using the carbohydrate recognition domain (CRD) and can be a powerful tool to study cancer glycan-based biomarkers.4 However, the real-time binding kinetics of these heterogenous glycan-lectin interactions have remained understudied due to the lack of efficient in vitro, label-free biophysical tools.3
In this application note, we study the interactions of glycans present in pancreatic cancer cells (BxPC3) with two different lectins, Wheat Germ Agglutinin (WGA) and Sambucus Nigra Lectin (SNA), using SPRm200, a Surface Plasmon Resonance Microscopy (SPRM). SPRm200 incorporates SPR and brightfield microscopy to detect binding of unlabeled lectin molecules to a heterogenous population of glycans in the native cell environment.
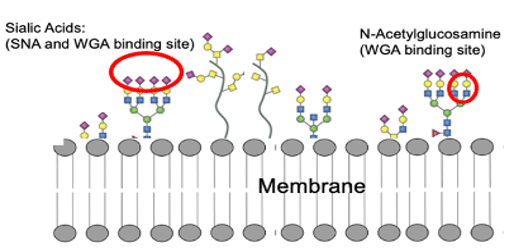
Figure 1: Schematic showing an example of aberrant surface glycosylation of pancreatic cancer cell and binding sites for two different lectins.
As shown in Figure 1, WGA binds to terminal N-acetyl-D-glucosamine and sialic acid, while SNA binds preferentially to sialic acid attached to terminal galactose in α-2,6 linkage and to a lesser degree, α-2,3 linkage.5&6 Even small changes in carbohydrate moieties may strongly affect interaction and consequent functionality. SPRM measures the kinetics of molecular interactions of ligands to cell-surface expressed receptors. Since SPRM does not require tagged or labeled ligands, it is beneficial for mitigating the potential adverse effects of labeling on ligand function and affinity.
In this study, SPRm 200 was used to validate the binding of the two different lectins to glycans on parental BxPC3 cells. To obtain binding kinetic parameters, all the kinetic interactions were quantified statistically by rendering the kinetic parameters into affinity isotherm plots and histograms, and by fitting the results with Gaussian distributions (Figure 2B and D). The kinetic parameters extracted from each responsive ROI from the cells were aggregated to form affinity isotherm plots using ImageSPR™analysis software (Figure 2A and C). The sub-cellular resolution of SPRM permit the usage of affinity isotherm plots which provide a uniquely informative representation of binding heterogeneity by revealing kinetic modes of binding interactions that would otherwise be hidden with traditional endpoint and affinity measurement techniques. The mean values of the distributions revealed the predominate values for the equilibrium dissociation (KD), association rate (ka), and dissociation rate (kd) constants.
Kinetic data assessing SNA and WGA binding to BxPC3 cells showed robust gaussian distributions for calculated on-rate, off-rate and binding affinity. The kinetic parameters for both the lectin binding interactions is shown in Table 1. WGA kinetic binding analysis to BxPC3 cells show two broad binding populations revealing greater glycan heterogeneity than SNA. These results support the currently held understanding that indeed SNA shows a much greater selectivity in the types of glycans that it binds to than those that WGA binds to.
| Lectin | ka x 105 [1/M*s] | kd x 10-4[1/s] | KD [nM] |
| SNA | 1.27 | 8.47 | 7.03 |
| WGA | 0.43 | 15.5 | 1661 |
| 4.64 | 366.1 | 42.13 |
Table 1: Summary of Lectin binding to BxPC3 pancreatic cells
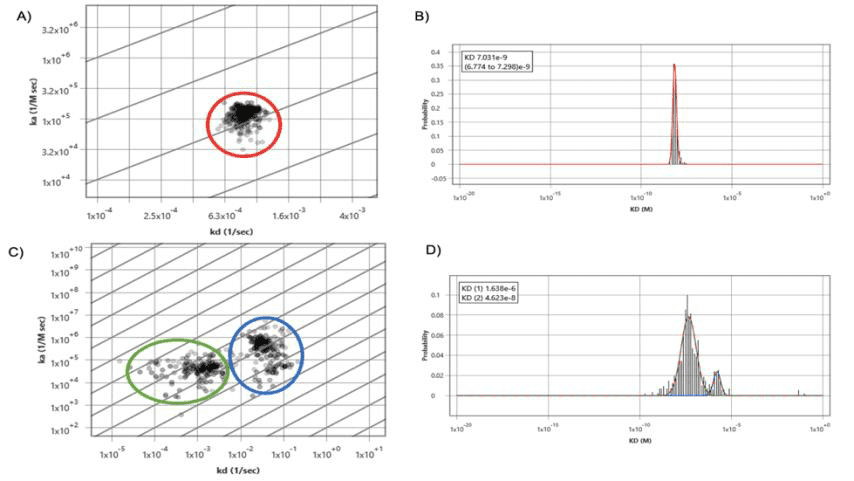
Figure 2: Kinetic interaction analysis was performed on BxPC3 cells using SPRm 200: (A) Affinity isotherm plot extracted from responsive ROIs showing one population highlighted in red of SNA lectin binding to glycans on BxPC3 cells. (B) Histogram of the kinetic parameters were fitted with a Gaussian distribution to extract the mean and 95% confidence interval of the cell population. (C) Affinity isotherm plot extracted from responsive ROIs showing WGA binding interactions with glycans in BxPC3 cells with two distinct populations highlighted in blue and green. (D) Histograms of the kinetic parameters were fitted with Gaussian distributions to extract the mean and 95% confidence interval of the cell populations.
These results indicate that SPRM is not only a useful tool to study lectin-glycan interaction but also to identify different modes of interaction depending on the target glycan residue. The presence of two populations in the isothermal plots for WGA binding indicate the presence of two different glycan binding sites (N-acetyl-D-glucosamine and sialic acid). These results highlight the complexity of lectin-glycan interactions in pancreatic cancer cells, and the need of a technology which monitors binding heterogeneity in real-time in whole cells. Furthermore, profiling the cell surface glycosylation using lectins not only plays a critical role in understanding the function of the complex glycocalyx in tumorigenesis, but could also be pivotal in the discovery of new diagnostics. This study is critical to understand fundamental biological processes and advance drug development by knowing more about the environment of different drug targets.
Author: Nguyen Ly, Miyuki Thirumurthy and Jesús Aguilar Díaz de león| Biosensing Instrument | Published June 10th, 2024
DOWNLOAD PDF
Download a PDF of Application Note 157: Understanding the Glyco-code using SPR microscopy
- Alberts B., Johnson A.D., Lewis J., Morgan D., Raff M., Roberts K., Walter P. Molecular Biology of the Cell. Garland Science; New York, NY, USA: 2014.
- Varki A., Kannagi R., Toole B., Stanley P. Essentials Glycobiology. 3rd ed. Cold Spring Harbor; New York, NY, USA: 2015. Glycosylation Changes in Cancer.
- Thomas, D., Rathinavel, A.K. and Radhakrishnan, P., 2021. Altered glycosylation in cancer: A promising target for biomarkers and therapeutics. Biochimica et Biophysica Acta (BBA)-Reviews on Cancer, 1875(1), p.188464.
- Hashim, Onn Haji, Jaime Jacqueline Jayapalan, and Cheng-Siang Lee. "Lectins: an effective tool for screening of potential cancer biomarkers." PeerJ 5 (2017): e3784.
- Kremsreiter, S.M., Kroell, A.S.H., Weinberger, K. and Boehm, H., 2021. Glycan–lectin interactions in cancer and viral infections and how to disrupt them. International Journal of Molecular Sciences, 22(19), p.10577.
- Cummings, R. D., Etzler, M., Hahn, M. G., Darvill, A., Godula, K., Woods, R. J., & Mahal, L. K. (2022). Glycan-recognizing probes as tools. Essentials of Glycobiology [Internet]. 4th edition
