The coronavirus 2019 (COVID-19) pandemic continues to evolve. This disease is caused by the highly contagious coronavirus 2 (SARS-CoV-2) and is initiated by the viral invasion into host cells through viral attachment to angiotensin (Ang)-converting enzyme 2 (ACE2) (1,2). ACE2, expressed in numerous different tissues, is the receptor through which the COVID-19 spike-protein (S-protein) gains entry into cells for subsequent viral replication. During viral infection, the trimeric S protein is cleaved into S1 and S2 subunits, where the S1 subunit contains the receptor binding domain (RBD), which directly binds to the peptidase domain of ACE2. Therefore, a better understanding of the binding interaction between S1 and S RBD proteins with ACE2 is important to the development of antiviral drugs.
Surface Plasmon Resonance (SPR) is widely used for kinetic measurements of molecular interactions. In this application note, we present the kinetic data between ACE2 and S1 protein binding measured using the BI-4500A system, a five channel SPR with the BI-DirectFlowTM technology. The immobilized ligand is a Biotinylated Human ACE2 with His-tag and Avi-tag (ACROBiosystems, AC2-H82E6), MW=87.2kDa. The analyte is SARS-CoV-2 S1 protein with Mouse IgG2a Fc Tag (ACROBiosystems, S1N- C5257) with MW=101kDa. The running buffer is 1xPBS (pH 7.2) + 0.05% Tween 20 + 0.1% BSA.
The biotinylated ACE2 protein was diluted in the running buffer and immobilized onto a streptavidin (SA) sensor chip using BI’s automated graduated injection technique, which immobilizes different amounts of ligand in each channel during a single injection. The ligand immobilization occurred in channel 1, 2, 3, and 4 for 200, 100, 50 and 10 s, respectively. Testing binding interactions with different amounts of immobilized ligand in each channel allows one to better examine any secondary effects due to ligand density.
S1 protein was prepared in the above running buffer. A series of binding kinetic titrations were performed with 3-fold analyte (S1 protein) dilution from 2 to 158 nM. A 45-min dissociation time was applied between consecutive injections. No regeneration was employed for this experiment. As expected, based upon the relative amount of immobilized ACE2 in each channel, channel 1 produced the largest binding responses and channel 4 produced the smallest binding responses (Figure 1).
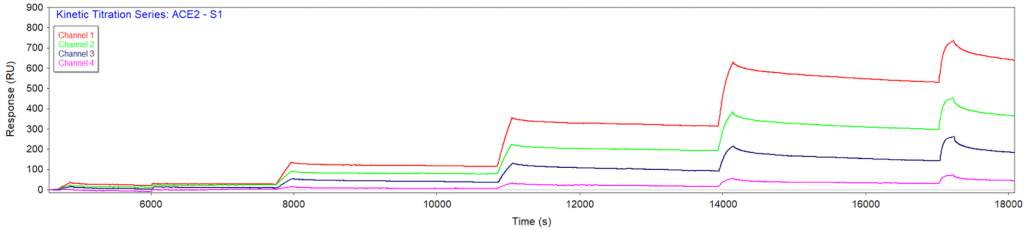
Figure 1 SPR binding response from each channel in a kinetic titration injection series of S1 protein. Each of the four channels had a different ACE2 immobilization amount (700, 400, 250 and 70 RU for channels 1-4, respectively) and all signals were measured simultaneously.
The experiment confirmed tight binding between the S1 and ACE2 proteins. Even after 45 min of dissociation, much of the S1 protein remained bound to the ACE2 protein. As a result, a multi-cycle kinetic-titration binding analysis was performed, where only the initial injection begins at zero, and successive injections have their fits extrapolated to zero. This approach enables simple and accurate analysis for non-regenerable interactions such as this one. The results of the kinetic analysis are presented in Figure 2. The measured binding responses closely overlay the kinetic fits in orange, and the extracted kinetic rate constants are in close agreement with literature, with KD ranging from hundreds of picomolar to tens of nanomolar. The binding parameters are summarized in the Table 1 below.
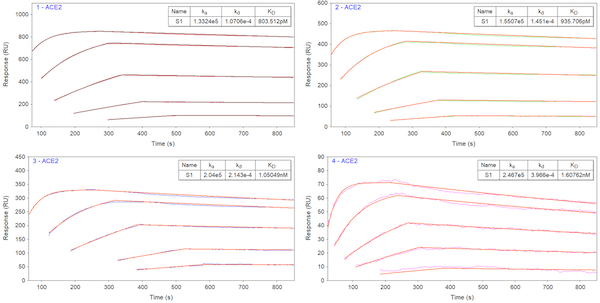
Figure 2 Kinetic binding analysis of S1 and ACE2 interaction. Sensorgrams are fitted with a kinetic-titration 1:1 interaction model. Kinetic binding interactions of all four channels are in close agreement with each other and with the literature value.
| Parameter | ka [1/M*s] | kd [1/s] | KD [nM] | *EC50 [nM] | **KD by BLI [nM] |
| Value | 1.85×105 | 2.15×10-4 | 1.10 | 5.99 | 3.95 |
* EC50 is 0.605 μg/m form FACS analysis of SARS-CoV-2 S1 protein, Mouse IgG2a Fc Tag (Cat. No. S1N-C5257) binding to Vero E6 cells surface ACE2.
** KD determined for SARS-CoV-2 S1 protein, Mouse IgG2a Fc Tag (Cat. No. S1N-C5257) binding to Human ACE2, His Tag (Cat. No. AC2-H52H8) in BLI assay.
This measurement using BI-4500A demonstrates the strong binding interaction of SARS-CoV-2 viral protein (S1) with ACE2 expressed on host cell surface, a key step of viral infection.
DOWNLOAD PDF
Download a PDF of Application Note 138: Kinetic Measurement of COVID-19 S1 Protein and ACE2 Binding
- Abassi Z, Higazi AAR, Kinaneh S, Armaly Z, Skorecki K and Heyman SN, ACE2, COVID-19 Infection, Inflammation, and Coagulopathy: Missing Pieces in the Puzzle. Front. Physiol. 11:574753 (2020)
- Renhong Yan, Yuanyuan Zhang, Yaning Li, Lu Xia, Yingying Guo, Qiang Zhou, Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2, Science 367, 1444–1448 (2020)
