Monoclonal antibody (mAb) therapies have become established methods for treating cancer, autoimmune disorders, asthma and many other diseases. MAb drugs represent approximately half of the total sales of all biopharmaceutical products (1) and over 60% of them target membrane proteins on the cell surface. (2) However, quantifying mAb-membrane protein binding affinity on single cells has been a difficult task.
Traditional SPR measures protein binding kinetics by immobilizing purified proteins onto a sensor surface. This approach is problematic for membrane proteins because they are difficult to extract from their cellular membrane environment and to keep their native conformations intact outside of their lipid environment.(3) SPRm 200 is a new label free technique that can measure the binding of antibodies to membrane proteins on cells without the need of extracting the proteins from the cell membranes. It also combines SPR with high-resolution optical microscopy to resolve and study single cells.
This application note describes the study of binding of two antibodies to membrane protein receptors on A172 cell homo sapiens brain glioblastom cells and H4 human brain neuroglioma cells, respectively. Each cell line was incubated on a sensor surface and fixed with 4% PFA (paraformaldehyde) at room temperature. Different concentrations of antibodies were injected into the SPRm 200 flow system and the affinity constant (KD) was derived from the Langmuir binding isotherm model. Kinetic titration method was used as the receptors were not regenerated.
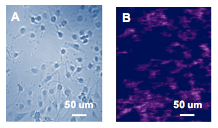
The A172 cells were first examined with bright field imaging capability of SPRm 200 to allow identification of individual cells (Fig. 1A). Antibody solutions of 8 different concentrations ranging from 1nM to 100 nM were then injected and the binding affinity of the antibody to membrane proteins on the identified cells were determined with SPRm 200 (Fig. 1B). The corresponding KD varies from cell to cell, ranging from 2.94 to 5.70 nM, which indicates the heterogeneity of the individual cells, and the average KD was found to be 4.3 nM.
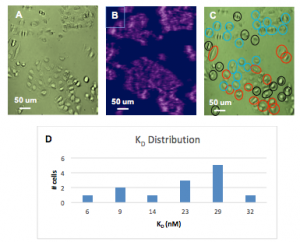
A similar study was performed for H4 cell line. Analysis of 44 individual H4 cells revealed variability of KD from1.4 to 31.2 nM with an average value of 16.8 nM (Fig. 2). The 44 cells can be divided into three ROI (region of interest) based on their phenotypic features shown in the bright field image (Fig. 2A). The KD of cells with distinct spherical shape (black ROIs, Fig. 2C) has a lower affinity (Fig. 2D) than other cell morphologies. This study provides unique information to understand how various cell types can be separated based on their different KD values.
In conclusion, SPRm 200 is a powerful technique for studying the binding kinetics of membrane proteins on cells without the need of extracting the membrane proteins from the cellular membranes. Its individual cell analysis capability allows study of heterogeneity of different cells.
We thank Prof. Takeshi Fukuhara, Dep. of Neurology, Juntendo University, Japan, for providing the cells and antibodies for this experiment.
Author: Nguyen Ly | Biosensing Instrument | Published Jan 4, 2025
DOWNLOAD PDF
Download a PDF of Application Note 127: Quantifying Antibody Binding to Membrane Proteins on Single Cells with SPR Microscopy
- Ecker, D et al, mAbs 7:1, 9-14 (2015)
- Jones, M et al, Scientific Reports, 6:26240 (2016)
- Wang, W et al, Nature Chemistry, 4, 846–853 (2012)