Endotoxin (commonly referred to as lipopolysaccharide in bacteriology) is associated with the outer membrane of Gram-negative bacterial pathogens such as Escherichia coli, Salmonella, Shigella, and Pseudomonas [1-2]. The interaction between endotoxin and bacterial cell surface is schematically depicted in Figure 1. Endotoxin elicits a series of pleiotropic effects on cells or organisms and is therefore harmful to most mammals. The most serious consequence of Gram-negative bacterial infection is caused by the circulation of endotoxin and is termed as ‘endotoxic shock’ [2]. An endotoxin molecule is less potent when it is membrane-bound. Thus pathological effects arise when endotoxin molecules are released from multiplying cells. It is known that endotoxin and its complexes with certain proteins in plasma can interact with the CD-14 protein, which is a type of receptors on monocytes, macrophages and endothelial cells. This application note describes a novel application of SPR to the detection of endotoxin via its interaction with the CD-14 protein.
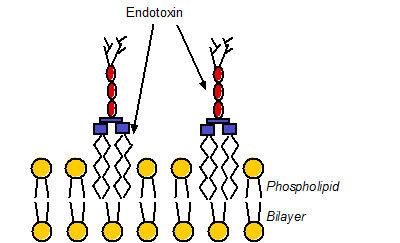
FIG. 1 The interaction of endotoxin with phospholipid bilayer of the outer membrane of Gram-negative bacterium. The yellow circles represent the lipid head groups and red ovals are the cores of endotoxin, and the branches are antigens. Endotoxin molecules are embedded in the bilayer via insertion of their lipid A chains into the membrane. For clarity, endotoxin and phospholipid molecules are not drawn to scale.
Before the experiment, a Au sensor chip was coated with a carboxymethylated dextran film. The formation of the carboxymethylated dextran is straightforward and follows a protocol described in literature [3]. Pyrogen-free water was used as the carrier (running) solution. Upon activation of the carboxyl groups on the carboxymethylated dextran with NHS/EDC, CD14 protein solution was injected into the SPR flow cell. CD14 can be effectively attached to the activated dextran film. Upon deactivation of the resultant surface with 1 M ethanolamine, solutions containing different amounts of endotoxin were injected and the binding curves were recorded. Figure 2 is an overlay of a series of binding curves.
Using a molecular weight of 10,000 Daltons for endotoxin, a KD value of 37 nM was deduced from the simulation and suggests that endotoxin binds strongly to CD-14 protein. Furthermore, the association (ka) and dissociation (kd) constants were determined to be 8.0 × 105 M-1 s-1 and 0.029 s-1, respectively. Thomas and Surolia measured the kinetics of the interaction between endotoxin and polymyxin B (a cationic decapeptide) and determined the KD value to be about 157 nM [4]. Comparable ka and kd values were also reported [4]. Thus, endotoxin appears to bind to CD14 almost as strongly as the peptide antibiotics. Moreover, an endotoxin concentration as low as 5 ng/mL can be easily detected. From this note, it is evident that SPR is a suitable technique for monitoring binding of endotoxins to proteins, antibiotics, and other species.
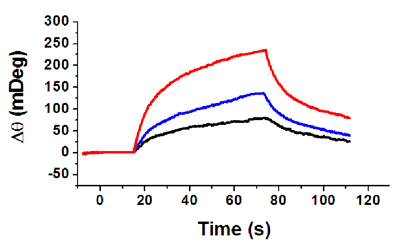
FIG. 2 Binding curves between pre-immobilized CD-14 protein and different concentrations of endotoxin: 20 ng/mL (red), 10 ng/mL (blue) and 5 ng/mL (black). A flow rate of 50 µL/min was used.
Author: Nguyen Ly | Biosensing Instrument | Published May 1, 2016
Acknowledgement: We thank Jim Powers for his technical help, discussion, and the provision of the CD-14 protein and endotoxin samples.
DOWNLOAD PDF
Download a PDF of Application Note: 109 – Application of SPR to Bacteriology: Endotoxin/Protein Interaction Studies
- Y.L. Tzeng, A. Datta, V.K. Kolli, R.W. Carlson and D.S. Stephens, J. Bacteriol. 184, 2379-88(2002)
- C.R.H. Raetz, Annu. Rev. Biochem. 59, 129-170(1990)
- X. Yao, X. Li, F. Toledo, C. Zurita-Lopez, M. Gutova, J. Momand and F. Zhou, Anal. Biochem. 354, 220-228(2006)
- C.J. Thomas and A. Surolia FEBS Lett. 445, 420-424(1999)
